HilyMax
Transfection Reagent
- Best Reagent for Cell Signaling Research
- High Transfection Efficiency in Wide Variety of Cells
- Optimized Protocol for Maximizing Transfection
- Great Result with Insect Cell
-
Product codeH357 HilyMax
| Unit size | Price | Item Code |
|---|---|---|
| 1 ml | Find your distributors | H357-10 |
| 1 ml x 5 | Find your distributors | H357-15 |
| 1 ml x 10 | Find your distributors | H357-20 |
| 1 ml | ・HilyMax Reagent ・Lipoform Buffer |
1 tube 1.0 ml×1 |
|---|---|---|
| 1 ml x 5 | ・HilyMax Reagent ・Lipoform Buffer |
5 tube 1.0 ml×5 |
| 1 ml x 10 | ・HilyMax Reagent ・Lipoform Buffer |
10 tube 1.0 ml×10 |
Description
Various methods have been developed to express specific proteins in mammalian cells. The first method to introduce DNA to cells was calcium phosphateprecipitation. However, the transfection efficiency was very poor and there was a high rate of cell-to-cell variation. The second method introduced was the DEAEsephadex method. The transfection efficiency drastically improved, but still the method could not be used for all cells and required heavy metalions to enhance transfection efficiency. The cation liposome method was then developed, which proved to be a much better method to transfect DNA and RNA into cells. Other methods used are magnet bead, metal particle shoot, and electroporation. However, the cationic liposome method does not require any special instruments or special skill. Therefore, many researchers are using this method. HilyMax is a newly developed gene transfection reagent that forms a liposome to be used for highly efficient gene transfection to a wide variety of cells. In addition, in signal transduction research, HilyMax gives better signal because the reagent introduced in cells does not interrupt intracellular signal pathways (Fig. 3). Since serum in the growth medium does not interfere with the transfection using HilyMax, no exchange of the medium during the transfection is required. HilyMax does not contain biological components that might interfere with the transfection.
Principle
.jpg)
HilyMax readily interacts with DNA because cationic liposome(+) and anionic DNA(-) spontaneously form DNAliposome complexes. The overall charge of DNA-HilyMax complex is positive, so that the DNA-HilyMax complex is electrostatically bound on an anionic cell surface and introduces DNA into the cell by endocytosis.
Procedure
.jpg)
The procedure is extremely simple. No exchange of the media is required during the whole process,
because serum in the medium does not interfere with the transfection.
Manual
Technical info
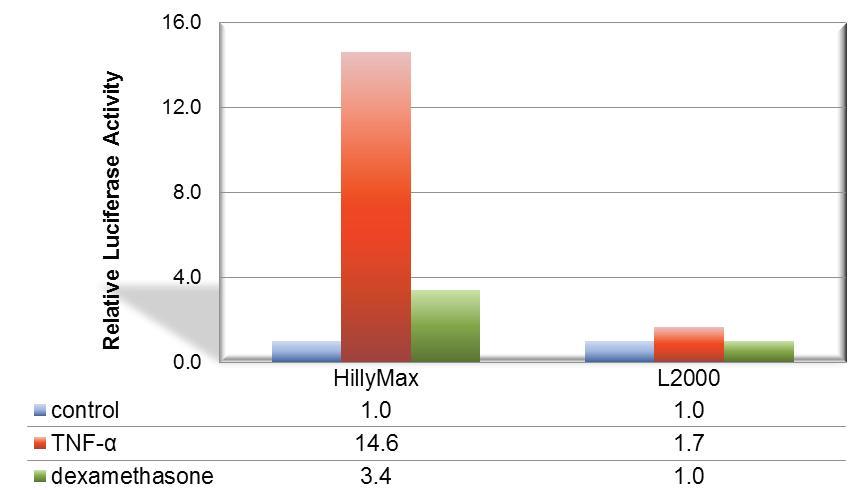
Figure 1. Suitable signal transduction research using HilyMax.
The signal transduction from A549 cell was confirmed with the TNF-a stimulation. For detection of cellular response, IL-8 dependent luciferase expression vector was transfected with HilyMax or L2000. The signal transduction response was detected as luciferase activity after stimulation and suppression. The signal response using HilyMax was related with the amount of expressed IL-8 in the stimulated cell.
If you are interested in this issue click here for more Information.
Comparison Data in Insect Cell
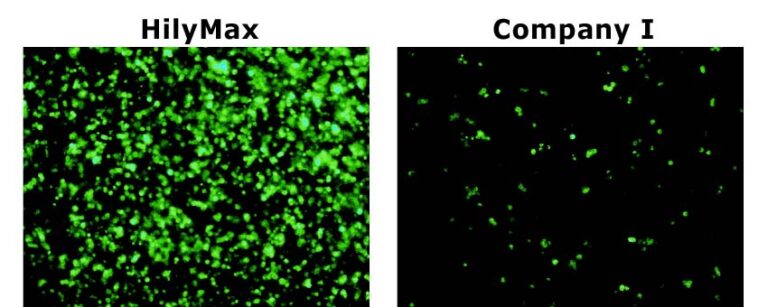
Data was kindly provided by Dr. Takashi Suzuki at Max Planck Institute of Neurobiology.
Culture Condition
Cell: S2(Schneider 2) cell, 200,000 cells/well
Media: Schneider’s Drosophila medium with 10% FCS
Antibiotics: 50 units Penicillin/ml, 20 µg Streptomycin/ml
Microplate: 24-well plate
Transfection Condition
Vector: 1 µg/well
Reagent: 5 µl/well
*Medium was changed in 4 hours after transfection.
*Schneider’s Drosophila medium(without seum and antibiotics) was used for the complex preparation.
GFP Transfected Cells with HilyMax
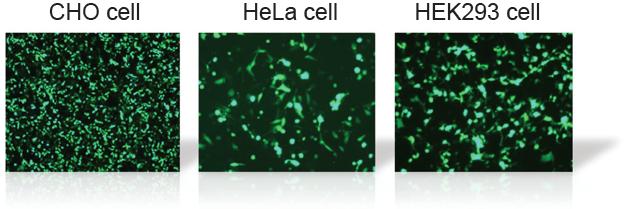
Transfection Efficiency in Various Cell Lines
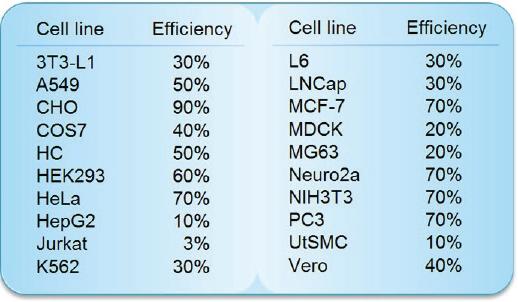
Comparison Data of Transfection Efficiency with Commercially Available Reagents
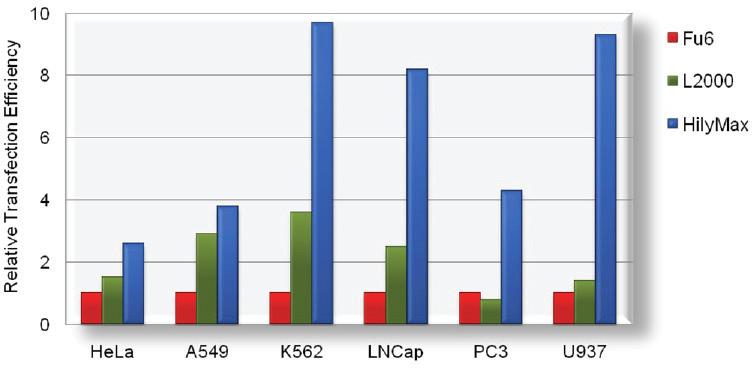
The tranfection efficiency of HilyMax is higher than commercially available reagents in widely used cells. GFP expressed DNA was transfected using HilyMax and the other transfection reagents in serum containing medium. The amount of expressed protein indicates the transfection efficiency.
Comprehensive Protocols for CRISPR/Cas9-Based Gene Editing Using HilyMax
Application of the CRISPR/Cas9 system to edit the genomes of human pluripotent stem cells (hPSCs) has the potential to revolutionize hPSC-based disease modeling, drug screening, and transplantation therapy.
Dr. Santos et.al. published protocols to enable groups —even those with limited experience with hPSC culture or the CRISPR/Cas9 system— to successfully perform genome editing using our HilyMax.
HilyMax is a useful tool to support generation of hPSC lines with gene-specific knock-outs, small targeted mutations, or knock-in reporters.
Reference
D. Santos, E. Kiskinis, K. Eggan, and F. Merkle, "Comprehensive protocols for CRISPR/Cas9-based gene editing in human pluripotent stem cells”, CurrProtoc Stem Cell Biol., 2016, 38, 5B.6.1–5B.6.60.
References
1) S. Mima, H. Ushijima, H.-J. Hwang, S. Tsutsumi, M. Makise, Y. Yamaguchi, T. Tsuchiya, H. Mizushima and T. Mizushima, "Identification of theTP01gene in Yeast, and its Human Orthologue TETRAN, which Cause Resistance to NSAIDs",FEBS Lett.,2007,581, 1457.
2) T. Namba, T. Ishihara, K. Tanaka, T. Hoshino and T. Mizushima, "Transcriptional Activation of ATF6 by Endoplasmic Reticulum Stressors",Biochem. Biophys. Res. Commun.,2007,355, 543.
3)H. Nakajima, W. Amano, A. Fujita, A. Fukuhara, Y. Azuma, F. Hata, T. Inui and T. Takeuchi, "The Active Site Cysteine of the Proapoptotic Protein Glyceraldehyde-3-phosphate Dehydrogenase Is Essential in Oxidative Stress-induced Aggregation and Cell Death",J. Biol. Chem.,2007,282(36), 26562.
4) A. Endo, D. Sumi and Y. Kumagai, "1,2-Naphthoquinone Disrupts the Function of cAMP Response Element-binding Protein through Covalent Modification",Biochem. Biophys. Res. Commun.,2007,36(1), 243.
Q & A
-
Q
Transfection efficiency is low.
-
A
1. The volume of HilyMax is not enough. Increase the HilyMax volume.
2. Cell density is too high. Reduce the cell density. The appropriate cell density for transfection is about 40-90%.
3. HilyMax reagent is not be dissolved completely. Please check that the HilyMax solution is homogeneous.
4. Incubation time for the preparation of HilyMax and DNA complex is too long.
5. Your culture medium for DNA-HilyMax complex formation contains serum and/or antibiotics. Please use serum-free and antibiotics-free medium for the complex formation.
-
Q
Toxicity is high. Most cells died.
-
A
1. Reduce the amount of DNA and/or HilyMax. Prepare the complex.
2. Cell density is too low. Reduce the cell density. The appropriate cell density for transfection is about 40-90%.
Handling and storage condition
| 0-5°C |