Biotin Labeling Kit - NH2 (for 1mg)
Protein Labeling Kit
-
Product codeLK55 Biotin Labeling Kit - NH2 (for 1mg)
| Unit size | Price | Item Code |
|---|---|---|
| 1 sample | $240.00 | LK55-10 |
| 1 sample | ・NH2-Reactive Biotin ・WS Buffer ・Reaction Buffer ・Filtration Tube ・15 ml Tube (for counterbalance) |
1 tube 13 ml x 1 1.2 ml x 1 1 tube 1 tube |
|---|
Description
Biotin Labeling Kit-NH2 is mainly used for the preparation of biotin-labeled IgG for enzyme immunoassay (EIA). NH2-reactive biotin, a component of this kit, has succinimidyl groups (NHS) that react with amino groups on proteins or other molecules (Fig. 1). This kit contains all the reagents necessary for the labeling. The labeling process is very simple. Just add the NH2- reactive Biotin to IgG solution and incubate at 37oC for 10 minutes. An average of 5 to 8 biotin molecules conjugate to each IgG molecule. The number of biotin molecules per protein can be determined by HABA assay. Excess biotin molecules can be removed by a filtration tube.
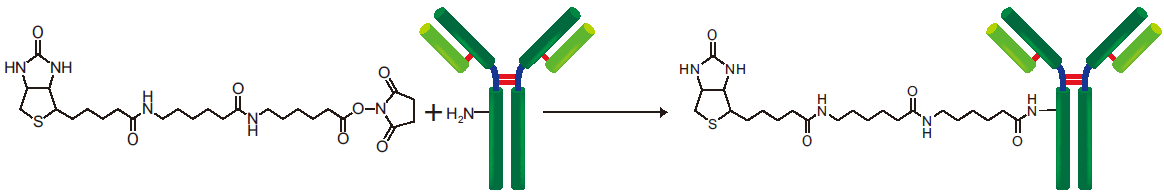
Fig. 1 IgG labeling reaction with NH2-reactive biotin
Precaution:
♦ The molecular weight of the protein to be labeled with this kit should be greater than 50,000.
♦ IgG or biotin-conjugated IgG is always on the membrane of the filtration tube during the labeling process.
♦ If the IgG solution contains other proteins with molecular weights larger than 10,000, such as BSA or gelatin, purify the IgG solution before labeling biotin with this kit.
♦ If the IgG solution contains small insoluble materials, centrifuge the solution and use the supernatant for the labeling.
Easily Switch Fluorescence wavelength on your primary antibody

| Developer | Dojindo Molecular Technologies, Inc |
|---|
Manual
Technical info
The average number of biotin molecules per IgG molecule should be in the range of 5 and 8. If you need to determine the precise number of biotin molecules per Protein molecule use HABA assay. The following is a HABA assay protocol.
Reagent solution:
200 μM HABA (4-hydroxyazobenzene-2-carboxylic acid) solution prepared with PBS, pH 7.4 …………………… 1 ml
0.5 mg avidin/ml solution prepared with PBS, pH 7.4 ………………………………………………………………….. 1 ml
diluted sample solution (55 μl biotinylated protein solution + 110 μl PBS, pH 7.4)
25 μM biotin prepared with a mixed solution (2 volumes of PBS, pH 7.4 + 1 volume of WS buffer)……………… 200 μl
Prepare solutions of various concentrations (12.5 μM, 6.25 μM, 3.13 μM, 1.56 μM) with serial dilution …………. 200 μl/ea
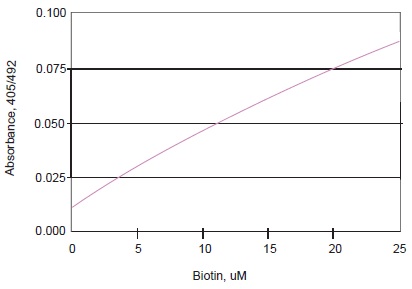
Fig. 2 Typical Calibration Curve of HABA Assay
1. Mix HABA solution and avidin solution in a plastic tube.
2. Add 100 μl of the HABA-avidin solution to 15 wells for multiple assays (n=3).
3. Add 50 μl biotin solution (12.5 μM, 6.25 μM, 3.13 μM, and 1.56 μM) to 3 wells each and 50 μl of diluted sample solution to the rest of the 3 wells.
4. Read the O.D. at 405 nm with a reference at 492 nm and prepare a calibration curve using the O.D. of various concentrations of biotin solution. Read the O.D. at 280 nm to determine the protein concentration. (e.g. molar absorptivity of IgG at 280 nm: 216,000).
5. Determine the concentration of biotin in the sample solution and calculate the number of biotin molecules per protein.
References
References
1. Y. Kubota, Y. Oike, S. Satoh, Y. Tabata, Y. Niikura, T. Morisada, M. Akao, T. Urano, Y. Ito, T. Miyamoto, N. Nagai, G. Y. Koh, S. Watanabe and T. Suda, “Cooperative Interaction of Angiopoietin-like Proteins 1 and 2 in Zebrafish Vascular Development”, Proc. Natl. Acad. Sci. USA, 2005, 102, 13502.
2. T. Yamabuki, A. Takano, S. Hayama, N. Ishikawa, T. Kato, M. Miyamoto, T. Ito, H. Ito, Y. Miyagi, H. Nakayama, M. Fujita, M. Hosokawa, E. Tsuchiya, N. Kohno, S. Kondo, Y. Nakamura, and Y. Daigo, “Dikkopf-1 as a Novel Serologic and Prognostic Biomarker for Lung and Esophageal Carcinomas”, Cancer Res., 2007, 67, 2517.
3. H. Kohara, Y. Omatsu, T. Suhiyama, M. Noda, N. Fujii and T. Nagasawa, “Development of Plasmacytoid Dendritic Cells in Bone Marrow Stromal Cell niches Requires CXCL12-CXCR4 Chemokine Signaling”, Blood, 2007, 110, 4153.
4. N. Ishikawa, A. Takano, W. Yasui, K. Inai, H. Nishimura, H. Ito, Y. Miyagi, H. Nakayama, M. Fujita, M. Hosokawa, E. Tsuchiya, N. Kohno, Y. Nakamura and Y. Daigo, “Cancer-testis Antigen Lymphocyte Antigen 6 Complex Locus K is a Serologic Biomarker and a Therapeutic Target for Lung and Esophageal Carcinomas”, Cancer Res., 2007, 67, 11601.
5. A. Fukuda, T. Goto, K.N. Kuroishi, K.K. Gunjigake, S. Kataoka, S. Kobayashi and K. Yamaguchi, “Hemokinin-1 competitively inhibits substance P-induced stimulation of osteoclast formation and function”, Neuropeptides., 2013, 47, (4), 251.
6. A. Ogawa , M. Sakatsume, X. Wang, Y. Sakamaki, Y. Tsubata, B. Alchi, T. Kuroda, H. Kawachi, I. Narita, F. Shimizu and F. Gejyo, “SM22alpha: the novel phenotype marker of injured glomerular epithelial cells in anti-glomerular basement membrane nephritis”, Nephron Exp. Nephrol.., 2007, 106, (3), e77.
7. D. Men, T.T. Zhang, L.W. Hou, J. Zhou, Z.P. Zhang, Y.Y. Shi, J.L. Zhang, Z.Q. Cui, J.Y. Deng, D.B. Wang and X.E. Zhang, “Self-Assembly of Ferritin Nanoparticles into an Enzyme Nanocomposite with Tunable Size for Ultrasensitive Immunoassay”, ACS Nano., 2015, 9, (11), 10852.
8. D. Sugahara, Y. Kobayashi, Y. Akimoto, H. Kawakami, “Mouse intestinal niche cells express a distinct α1,2-fucosylated glycan recognized by a lectin from Burkholderia cenocepacia”, Glycobiology., 2017, 27, (3), 246.
9. H. Sasaki-Iwaoka, M. Ohori, A. Imasato, K. Taguchi, K. Minoura, T. Saito, K. Kushima, E. Imamura, S. Kubo, S. Furukawa and T. Morokata, “Generation and characterization of a potent fully human monoclonal antibody against the interleukin-23 receptor”, Eur. J. Pharmacol.., 2018, 828, 89.
10. K. Kaneshiro, M. Watanabe, K. Terasawa, H. Uchimura, Y. Fukuyama, S. Iwamoto, T.A. Sato, K. Shimizu, G. Tsujimoto and K. Tanaka, “Rapid quantitative profiling of N-glycan by the glycan-labeling method using 3-aminoquinoline/α-cyano-4-hydroxycinnamic acid”, Anal. Chem.., 2012, 84, (16), 7146.
11. K.S. Tan, K. Kulkeaw, Y. Nakanishi, D. Sugiyama, “Expression of cytokine and extracellular matrix mRNAs in fetal hepatic stellate cells”, Genes Cells., 2017, 22, (9), 836.
12. M. Tahara, S. Ohno, K. Sakai, Y. Ito, H. Fukuhara, K. Komase, M.A. Brindley, P.A. Rota, R.K. Plemper, K. Maenaka and M. Takeda, “The receptor-binding site of the measles virus hemagglutinin protein itself constitutes a conserved neutralizing epitope”, J. Virol.., 2013, 87, (6), 3583.
13. M. Takahashi, Y. Ishida, D. Iwaki, K. Kanno, T. Suzuki, Y. Endo, Y. Homma and T. Fujita, “Essential role of Mannose-binding lectin-associated serine protease-1 in activation of the complement factor D”, J. Exp. Med.., 2010, 207, (1), 29.
14. N. Hosen, Y. Matsunaga, K. Hasegawa, H. Matsuno, Y. Nakamura, M. Makita, K. Watanabe, M. Yoshida, K. Satoh, S. Morimoto, F. Fujiki, H. Nakajima, J. Nakata, S. Nishida, A. Tsuboi, Y. Oka, M. Manabe, H. Ichihara, Y. Aoyama, A. Mugitani , T. Nakao, M. Hino, R. Uchibori, K. Ozawa, Y. Baba, S. Terakura, N. Wada, E. Morii, J. Nishimura, K. Takeda, Y. Oji, H. Sugiyama, J. Takagi and A. Kumanogoh, “The activated conformation of integrin β7 is a novel multiple myeloma-specific target for CAR T cell therapy”, Nat. Med.., 2017, 23, (12), 1436.
15. R. Nishino, A. Takano, H. Oshita, N. Ishikawa, H. Akiyama, H. Ito, H. Nakayama, Y. Miyagi, E. Tsuchiya, N. Kohno, Y. Nakamura and Y. Daigo, “Identification of Epstein-Barr virus–induced gene 3 as a novel serum and tissue biomarker and a therapeutic target for lung cancer”, Clin. Cancer Res.., 2011, 17, (19), 6272.
16. T. Hiono, A. Matsuda, T. Wagatsuma, M. Okamatsu, Y. Sakoda and A. Kuno, “Lectin microarray analyses reveal host cell-specific glycan profiles of the hemagglutinins of influenza A viruses”, Virology., 2019, 527, 132.
17. T. Oshima, S. Sato, J. Kato, Y. Ito, T. Watanabe, I. Tsuji, A. Hori, T. Kurokawa and T. Kokubo, “Nectin-2 is a potential target for antibody therapy of breast and ovarian cancers”, Mol. Cancer., 2013, 12, (60), .
18. T. Yoshida, N. Shiraki, H. Baba, M. Goto, S. Fujiwara, K. Kume and S. Kume, “Expression patterns of epiplakin1 in pancreas, pancreatic cancer and regenerating pancreas”, Genes Cells., 2008, 13, (7), 667.
19. X. Piao, T. Ozawa, H. Hamana, K. Shitaoka, A. Jin, H. Kishi and A. Muraguchi, “TRAIL-receptor 1 IgM antibodies strongly induce apoptosis in human cancer cells in vitro and in vivo”, Oncoimmunology., 2016, 5, (5), e1131380.
20. Y. Shimazaki, Y. Kohno, “Successive analysis of antigen trapping and enzymatic digestion on membrane-immobilized avidin”, Anal. Biochem.., 2012, 422, (1), 55.
Q & A
-
Q
Can I use this kit for other proteins?
-
A
Yes, if the molecular weight is greater than 50,000.
-
Q
Do I have to use a Filtration tube prior to labeling the protein?
-
A
If the protein solution does not contain small molecules with amino groups and the concentration of the protein is 10 mg per ml or about 70 μM, there is no need to use the Filtration tube. Just mix 10 μl of the sample solution with 90 μl of Reaction buffer and add the mixture to a vial of NH2-reactive Biotin. After the reaction, transfer all of the reaction mixture to a Filtration tube, and then follow the protocol starting at step 6.
-
Q
Do I have to use WS buffer to store the biotin-labeled protein?
-
A
You don’t have to use WS buffer. You can choose any kind of buffer according to your experiment.
-
Q
My sample contains small insoluble material. What should I do?
-
A
Spin the sample and use the supernatant for the labeling.
-
Q
How long is the biotin-labeled protein stable?
-
A
If you store the biotin-labeled protein at 0-5ºC, it is stable for 2 months. For longer storage, add 100% volume of glycerol, aliquot, and store at -20ºC (if the protein can be frozen). However, please note that stability depends on the protein itself.
-
Q
What is the minimum amount of IgG that can be labeled by this kit?
-
A
The minimum amount is 10 μg IgG; simply follow the protocol. The labeling ratio remains the same for 10 μg to 100 μg of IgG.
Handling and storage condition
| 0-5°C, Protect from moisture |