|
Expansion microscopy is a useful method that physically expands gel-anchored specimens several-fold, thereby improving the effective spatial resolution of fluorescence imaging and enabling analysis of intracellular nanostructures. At the same time, challenges remain in molecular retention during sample processing and in achieving high-contrast imaging, and various efforts have been made to broaden the scope of ExM. Recently, progress has been reported in two directions. One study introduced Land-ExM, which enabled high-contrast retention of protein and lipid signals together with 3D visualization of organelles and contact sites in combination with immunostaining. Another study developed a fixable and clickable cholesterol probe, enabling nanoscale imaging of cholesterol together with organelle markers. Together, these studies advance ExM as a platform for integrating intracellular molecular distribution with ultrastructural context. |
||||||||||
|
Landscape expansion microscopy reveals interactions between membrane and phase-separated organelles (Journal of Cell Biology, 2026) Highlighted technique: The authors designed land-ExM around NHS-biotin-MA, a trifunctional molecule that reacts with primary amines, covalently anchors labeled components to the hydrogel, and enables post-expansion fluorescent detection through biotin–streptavidin binding. Cellls were fixed and labeled with mCLING for lipids, followed by optional immunostaining, and NHS-biotin-MA was then used to co-anchor proteins and the lipid probe for multichannel ExM imaging. |
||||||||||
|
A Multifunctional Probe for Visualization of the Nanoscale Distribution of Cholesterol in Cells by Expansion Microscopy (Aggregate, 2025) Highlighted technique: Expansion microscopy has advanced nanoscale imaging in cells, but applying it to cholesterol has been difficult because cholesterol is poorly retained during fixation and easily lost during later processing. In this study, the authors expand the scope of ExM by developing a fixable and clickable cholesterol probe, enabling nanoscale visualization of cholesterol together with organelle markers and providing a foundation for extending ExM to other chemically challenging lipid-like small molecules. |
||||||||||
| Related Techniques | ||||||||||
| All-in-one reagents for expansion microscopy | Three-Dimensional Expansion Microscopy Kit (4x 3D-ExM) | |||||||||
| Nuclear staining | DAPI, Hoechst 33342 and Hoechst 33258 | |||||||||
| Related Applications | ||||||||||
Application Note (click to open/close)
|
||||||||||
|
After staining the nuclei and tubulin of RPE1 cells, the cells were expanded using this kit. The samples after expansion were imaged by confocal microscopy with a 60× oil-immersion objective, and 3D imaging was performed. As a result, the detailed morphology of the nuclear structures and cytoskeleton could be observed three-dimensionally at high resolution (Movie 1). Furthermore, two nuclei with different shapes present within the same cell could be compared and their structural differences could be confirmed in three dimensions (Movie 2). Experimental data:Kindly provided by Dr. Aussie Suzuki, University of Wisconsin–Madison.
|
|||||
> Mitochondrial Morphological Changes Induced by Different Drug Treatments
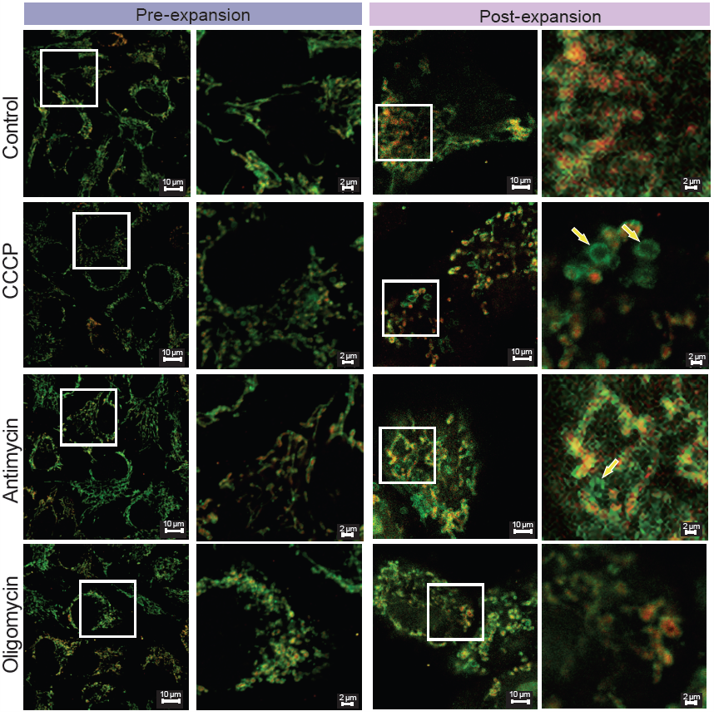 |
<Experimental Conditions> |















