Kit Contents
| Monomer Solution | 3 ml × 1 |
| Polymerization Initiator | 30 mg × 1 |
| Polymerization Promoter | 250 µl × 1 |
| Activation Buffer | 3 ml × 1 |
| Enzyme Solution | 100 µl × 1 |
| Crosslinking Solution | 100 µl × 1 |
| Mold | × 10 |
| Cover Glass | × 10 |
Storage Conditions
Store at 0–5 ℃
Required Equipment and Materials
- 2–20 µl, 20–200 µl, 100–1000 µl micropipettes
- 0.5 ml, 1.5 ml microtubes
- 18G needle
- Tweezers
- Microscope slides
- Humidified chamber for gel preparation (such as a 1 ml pipette-tip case)
- 10 cm dish
- Humidified chamber for expansion (such as a plastic food container with a moistened paper towel)
- Razor blade or scalpel
- PBS (–)
- DAPI solution (product code: D523)
- 24-well plate
- Spatula
- Graduated cylinder
- Ice bath
General Information
Multiple organelles exist within cells, and visualizing their structures is essential to understand cellular functions and disease mechanisms. High-resolution imaging at the organelle level typically requires expensive electron microscopes or super-resolution microscopes. In contrast, the resolution of conventional fluorescence microscopes is limited by Abbe’s diffraction limit, which makes structural analysis challenging. Expansion Microscopy (ExM), developed in 2015, enables physical magnification of specimens using a water-swellable polymer gel, allowing fluorescence microscopes to image fine structures that are otherwise difficult to resolve.
This kit contains the reagents and the mold required for gel preparation of ExM, enabling approximately 4-fold three-dimensional expansion of cells.
Precautions
- Bring all kit reagents to room temperature before use.
- Store the gel under water or in a Humidified chamber to prevent drying.
- Handle Cover Glass with care to avoid breakage.
Preparation of Solutions and Pre-treatment
Preparation of Polymerization initiator solution
- 1. Add 270 µl of ddH₂O to the Polymerization Initiator in a microtube and dissolve.
- 2. Aliquot Polymerization initiator solution into 0.5 ml microtubes, 25 µl each, and store at −20 °C.
- After preparation, store immediately at −20 °C. The prepared solution is stable for one month.
Preparation of Digestion solution
Add 223 µl of Activation Buffer and 7 µl of Enzyme Solution to a 1.5 ml microtube, then mix by pipetting.
- The Digestion Solution cannot be stored. Use it with in the day of preparation.
Preparation of Crosslinking working solution
Add 223 µl of PBS (–) and 7 µl of Crosslinking Solution to a 1.5 ml microtube, then vortex to mix.
- The Crosslinking working solution cannot be stored. Use it with in the day of preparation.
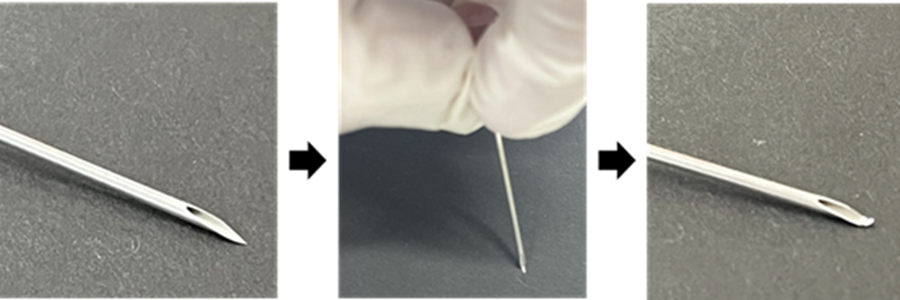
Preparation of a bent-tip needle
Carefully press the tip of an 18G needle against a flat surface to bend it.
- Use caution to avoid injury.
 |
General Protocol
- Place a Cover Glass into a well of a 24-well plate using tweezers.
- Perform cell culture on a clean bench.
- Seed cells onto the Cover Glass and incubate overnight in an incubator (37 °C, 5 % CO₂).
- Perform the desired staining and nuclear staining with 10 µg/ml DAPI aqueous solution.
- Nuclear staining makes it easier to find cells under the microscope.
- Add 200 µl of Crosslinking working solution to each well, protect from light (e.g., wrap in aluminum foil), and shake gently on a shaker at room temperature overnight.
- Remove the supernatant and wash cells 5 times with 200 µl of PBS (–). After washing, add 200 µl of PBS (–) to each well.
- Peel off the double-sided protective films from the Mold, and attach the Mold to a slide glass.
- Ensure no air is trapped between the Mold and the slide to prevent leakage of the solutions.
- Raise the Cover Glass (step 5) with a bent-tip needle. Pinch and transfer it with tweezers into the circular area of the Mold (step 6).
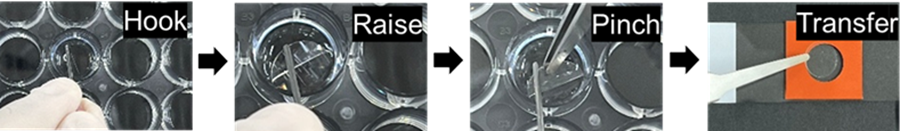

(Operation video: https://www.youtube.com/embed/LpseTF43aE0) - Add 100 µl of Monomer Solution onto the Cover Glass and incubate at 4 °C for 30 min.
- It is acceptable if the Monomer Solution spreads to fill the entire circular area inside the Mold.
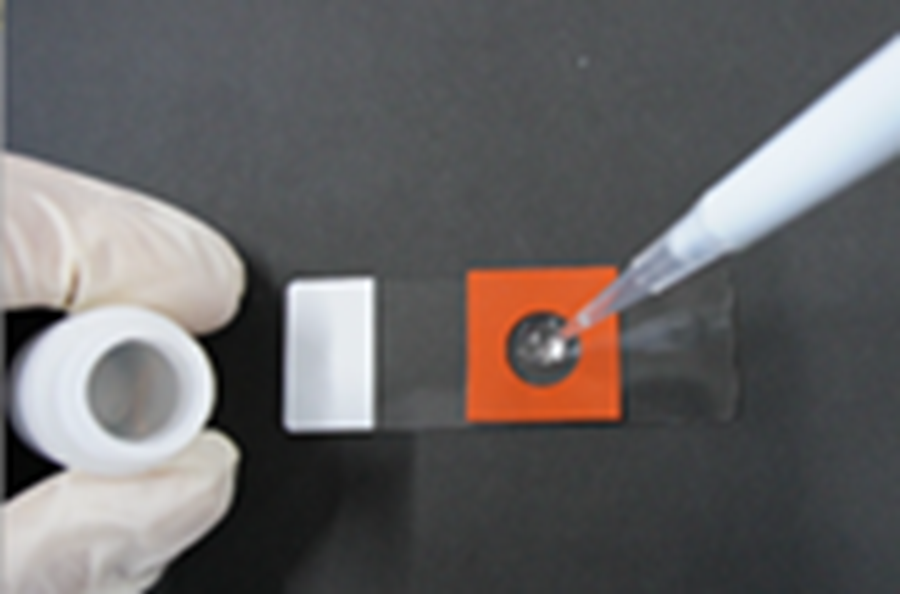

- Prepare the humidified chamber for gelation during step 8:
Place a wet paper towel at the bottom of a 1 ml pipette-tip case, close the lid, and incubate at 37 °C in an incubator.

- Aspirate the solution (step 8), then add 100 µl of the gelling solution to the Cover Glass.
Prepare the Gelling Solution as described below.
Add 190 µl of Monomer Solution, 2 µl of ddH₂O, and 4 µl of Polymerization Promoter to a 1.5 ml microtube on ice. After vortexing, add 4 µl of Polymerization initiator solution and mix by pipetting.
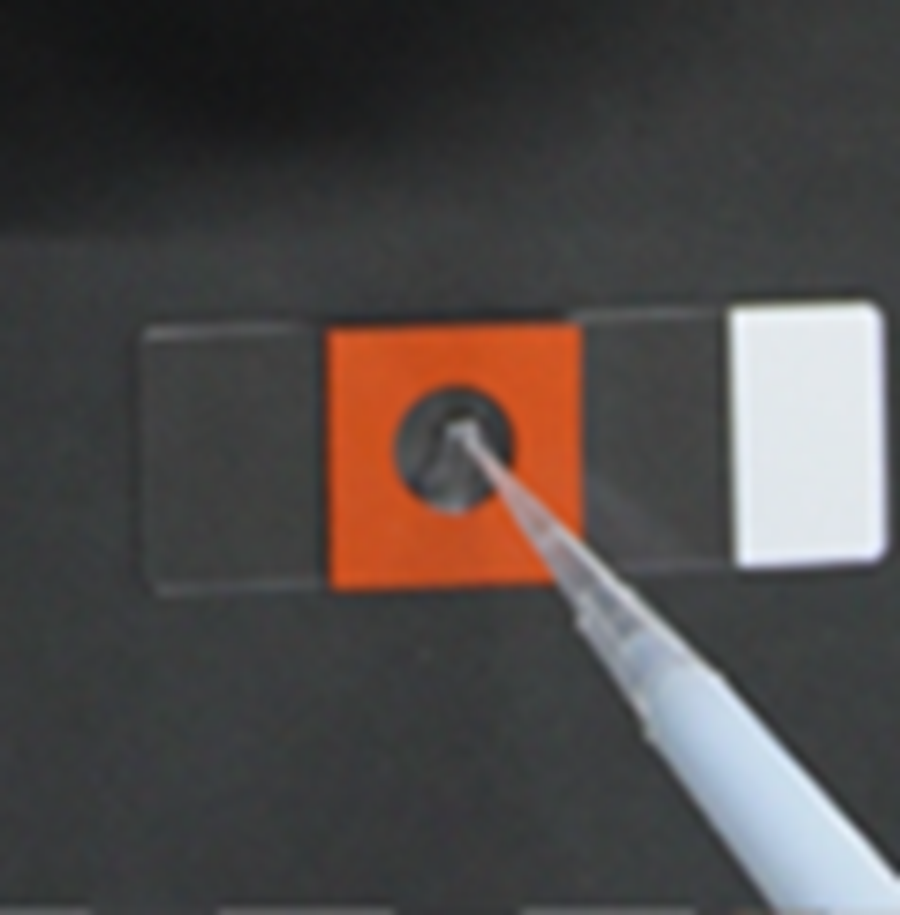

- Gelation starts upon addition of the Polymerization Initiator Solution. Add the Polymerization Initiator Solution immediately before Step 10, “Add 100 µl of the Gelling Solution onto the cover glass,” and use the solution within 10 minutes after preparation.
- Place the slide (step 10) into the pre-warmed humidified chamber (step 9), close the lid, and incubate at 37 °C for 1 h.
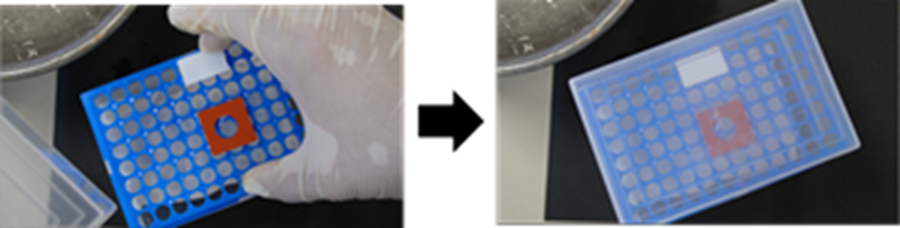

- Pull the chamber out of the incubator, then add 200 µl of the Digestion solution into the circular area of the Mold. After closing the lid, incubate at 37 °C for 3 h.
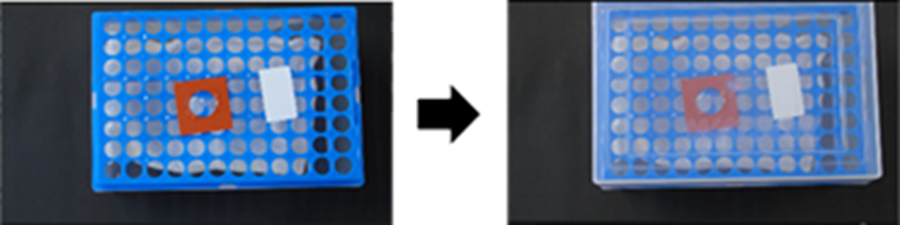

- Prepare the humidified chamber for expansion during step 12:
Use a container that can hold a 10 cm dish.
Place a wetted paper towel on the bottom, close the lid, and incubate at 37 °C.


- Add about 45 ml of ddH₂O to a 10 cm dish.
- Pull the chamber out of the incubator, and take out the slide (step 12) from the chamber.
- Detach the Mold gently from the slide, and transfer the gel into the 10 cm dish (step 14).
- Minor scratches at the gel edge from removing the Mold are acceptable.
- It is acceptable to place the Cover Glass into the 10 cm dish with the gel.
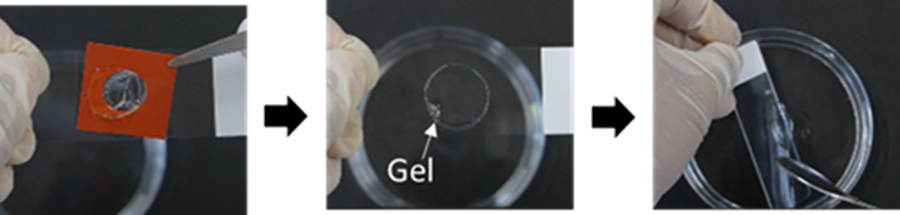

(Operation video:https://www.youtube.com/embed/bzpiKpsb9LE) - Gently swirl the 10 cm dish in a circular motion to ensure the gel is fully immersed in ddH₂O.
- Place the 10 cm dish (without its lid) in the warmed humidified chamber (step 13) and incubate at 37 °C for 30 min.
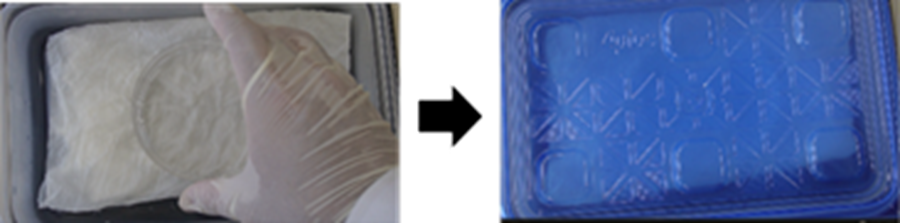

- Carefully aspirate all water from the 10 cm dish, avoiding damage to the gel.
- Use an aspirator, micropipette, or dropper.

(Operation video:https://www.youtube.com/embed/959AFnx8HHw) - Add approximately 60 ml of ddH₂O to the 10 cm dish, and incubate at 37 °C for 30 min.
- Add ddH₂O carefully to avoid damaging the gel.

- Repeat steps 19 and 20 twice more.
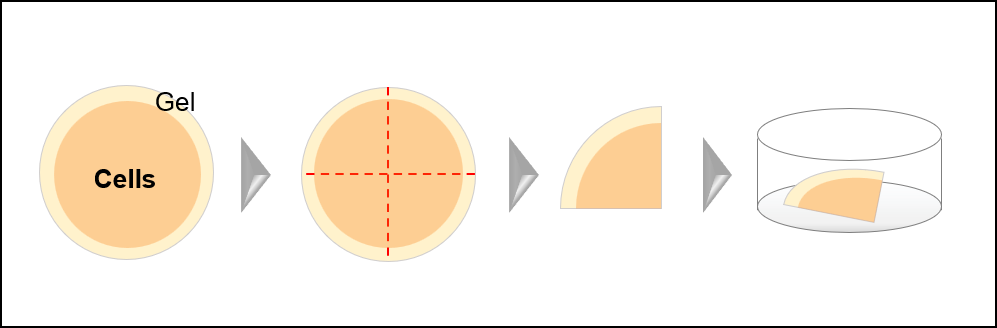
- Trim the expanded gel as shown in the figure below.
- Using a spatula, gently transfer the gel to a 35 mm observation dish without damaging it.
- Acquire images on a fluorescence microscope.
- Please check the following points before observation.
For detailed imaging procedures, please refer to the "Imaging Manual."
- A water-immersion objective lens is recommended.
- Cells are difficult to observe in bright-field or phase-contrast microscopy. Focus on the DAPI-stained nuclei.
- If the fluorescence intensity of DAPI-stained nuclei is low, add 3 ml of 10 µg/ml DAPI aqueous solution to the 35 mm dish and incubate at room temperature for 30 min.After incubation, wash the gel with ddH₂O and proceed to imaging.
- If observation is difficult, the gel may be upside-down. In that case, turn the gel over with a spatula and observe again.
- If the fluorescence intensity is low, add PBS (–) to the 35 mm dish and incubate for 15–30 min. Perform additional staining (such as secondary immunostaining). After staining, wash the gel 3 times with ddH₂O and re-expand it.
- Please check the following points before observation.
Experimental Example
Imaging of the mitochondrial outer and inner membranes in uncoupler-treated HeLa cells
[Cell Preparation]
- A Cover Glass was placed in a 24-well plate. HeLa cells (2.0 × 10⁵ cells/mL, 500 µl) in MEM (10% FBS) were seeded onto the Cover Glass and cultured overnight at 37 °C in a 5 % CO₂ incubator.
- After removing the supernatant, the cells were washed once with 500 μl of HBSS (−).
[Drug Treatment] - CCCP (5 µmol/l or 10 µmol/l) in MEM (serum free) was added to each well (200 µl), and the cells were incubated at 37 °C for 30 min.
[Fixation and Staining] - Paraformaldehyde (4% w/v) in PBS (200 μl) was added, and the cells were incubated at room temperature for 15 min.
- After removing the supernatant, the cells were washed three times with 200 μl of PBS.
- NH₄Cl (0.1 mol/l) in PBS (200 μl) was added, and the cells were incubated at room temperature for 10 min to quench residual aldehydes.
- After removing the supernatant, the cells were washed three times with 200 μl of PBS.
- Triton X-100 (0.1% v/v) in PBS (200 μl) was added, and the cells were incubated at room temperature for 10 min.
- After removing the supernatant, the cells were washed three times with 200 μl of PBS.
- Blocking One-P (nacalai tesque 05999-84) (10% v/v) in PBS (200 μl) was added, and the cells were incubated at room temperature for 1 h.
- After removing the supernatant, anti-COX IV (rabbit monoclonal IgG) and anti-TOM20 (mouse monoclonal IgG) antibodies diluted 1:200 in Can Get Signal Immunoreaction Enhancer Solution (TOYOBO NKB-101) (200 μl) were added and the cells were incubated at room temperature for 2 h.
- After removing the supernatant, the cells were washed three times with 200 μl of PBS.
- After removing the supernatant, Alexa Fluor 568-labeled anti-rabbit IgG and Alexa Fluor 488-labeled anti-mouse IgG diluted 1:200 in Can Get Signal Immunoreaction Enhancer Solution (200 μl) were added and the cells were incubated at room temperature for 1 h in the dark.
- After removing the supernatant, the cells were washed three times with 200 μl of PBS.
- 1 mg/ml DAPI solution diluted 1:100 in PBS (200 µl) was added, and the cells were incubated at room temperature for 30 min.
- After removing the supernatant, the cells were washed three times with 200 μl of PBS.
[Crosslinking and Gelation] - Crosslinking working solution (200 µl) was added to each well. The plate was covered with aluminum foil to protect it from light, and gently shaken on a shaker at room temperature, overnight.
- After removing the supernatant, the cells were washed five times with 200 μl of PBS, and 200 µl of PBS (−) was added to each well.
- The Mold was attached to a slide glass.
- The Cover Glass, after the staining procedures was raised with a bent-tip needle, and transferred into the circular area of the Mold with tweezers.
- Monomer Solution (100 µl) was added onto the Cover Glass, and incubated at 4 °C for 30 min.
- After removing the supernatant, Gelling solution (100 µl) was added onto the Cover Glass. The slide was placed into a pre-warmed humidified chamber, and incubated at 37 °C for 1 h.
[Digestion and Expansion] - Digestion solution (200 µl) was added into the circular area of the Mold and incubated at 37 °C for 3 h.
- After detaching the Mold gently from the slide, the gel was transferred into the 10 cm dish containing 45 ml of ddH₂O. The dish was placed in the warmed humidified chamber, and incubated at 37 °C for 30 min.
- All the solution was carefully aspirated from the dish, taking care not to damage the gel. Approximately 60 ml of ddH₂O was added to the dish, and incubated at 37 °C for 30 min.
- Step 25 was repeated twice more.
[Imaging] - All the solution was carefully aspirated from the dish, taking care not to damage the gel. The expanded gel was trimmed to about 2 cm × 2 cm, and gently transferred to a µ-Dish 35 mm, High Glass Bottom (ibidi cat. no. 81158) using a spatula.
- The gel was observed under a confocal laser scanning microscope (LSM 800, Zeiss).
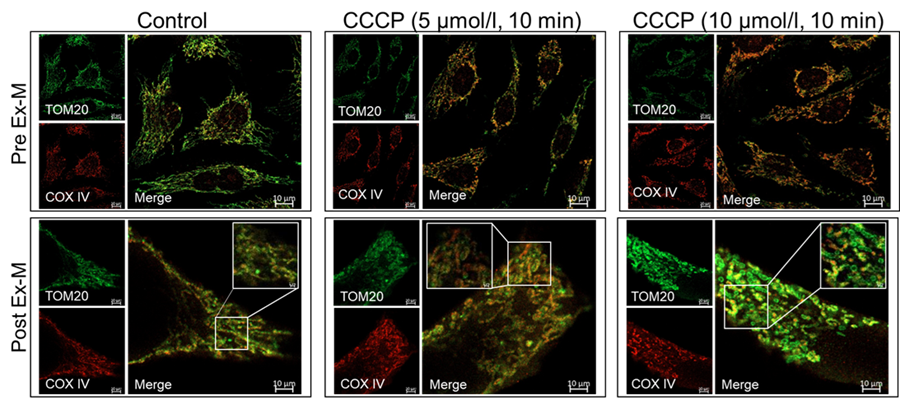
Pre Ex-M
TOM20) Laser Power:1%, Digital Gain:650V Excitation:488 nm Emission:500-550 nm
COXⅣ) Laser Power:2%, Digital Gain:650V Excitation:561 nm Emission:580-620 nmPost Ex-M
TOM20) Laser Power:70%, Digital Gain:700V Excitation:488 nm Emission:500-550 nm
COXⅣ) Laser Power:70%, Digital Gain:750V Excitation:561 nm Emission:580-620 nm
Figure1 Morphological changes in mitochondria subjected to drug treatment before and after Expansion Microscopy
Reference
1) R. X. Norman et al., ournal of Cell Biology, 2025, 224, e202407116.
This product is covered by U.S. Patent Application No. 17/466,434.
This technology is based on research conducted by Dr. Aussie Suzuki at the University of Wisconsin–Madison.
Frequently Asked Questions / Reference
T517: Three-Dimensional Expansion Microscopy Kit (4x 3D-ExM)
Revised Feb., 04, 2026


 Hidden sections will not be printed.
Hidden sections will not be printed.

